Properly Setting the Random Seed in ML Experiments. Not as Simple as You Might Imagine | by ODSC - Open Data Science | Medium

Accelerating Inference in TensorFlow with TensorRT User Guide :: NVIDIA Deep Learning Frameworks Documentation
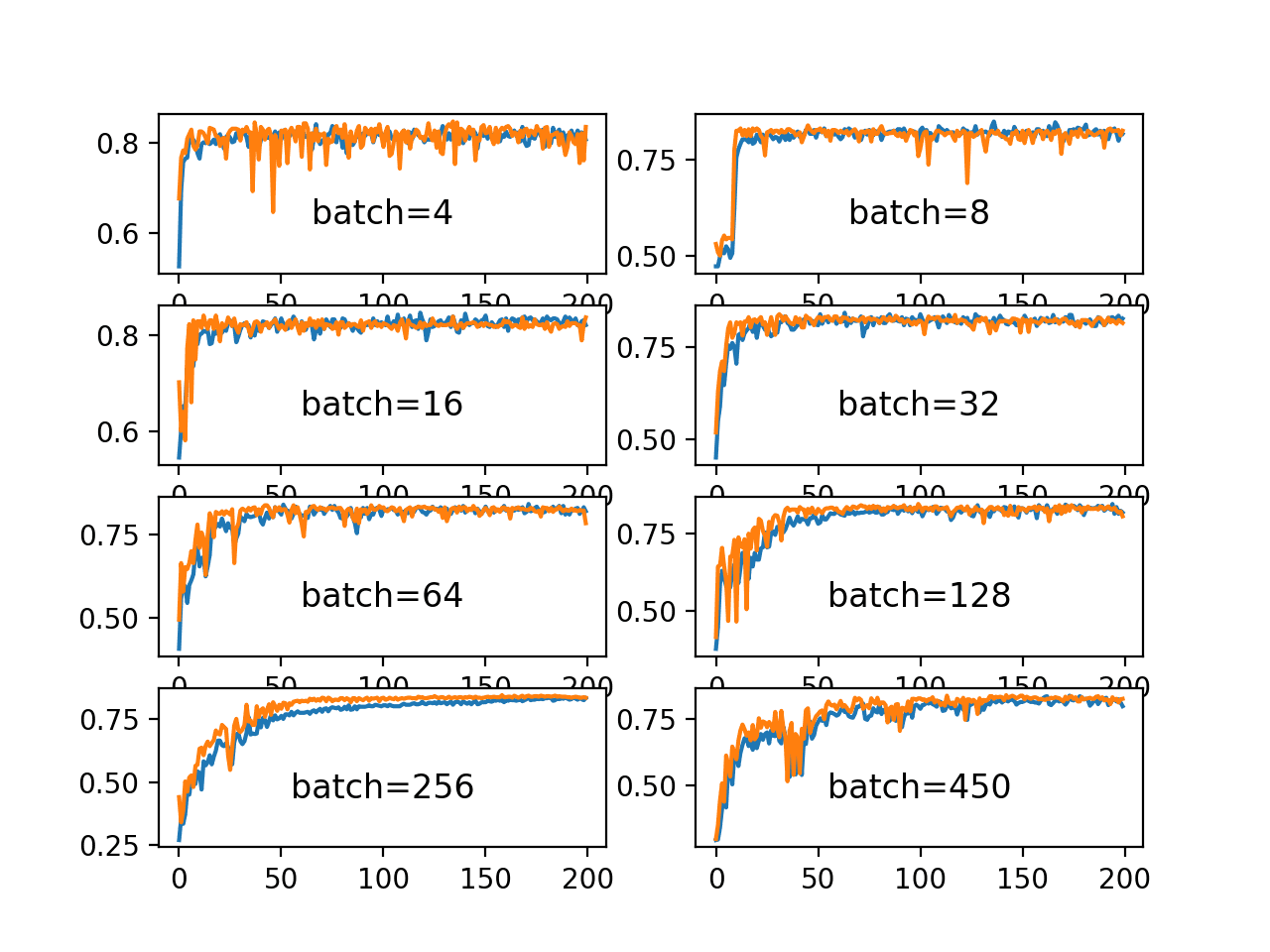
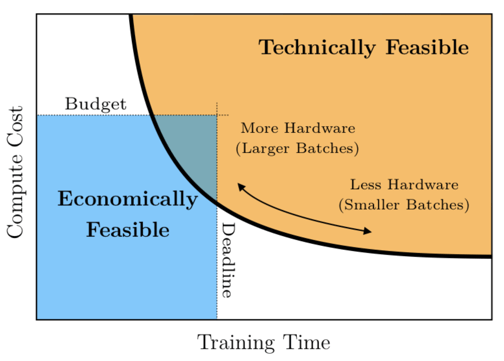
How to Control the Stability of Training Neural Networks With the Batch Size - MachineLearningMastery.com
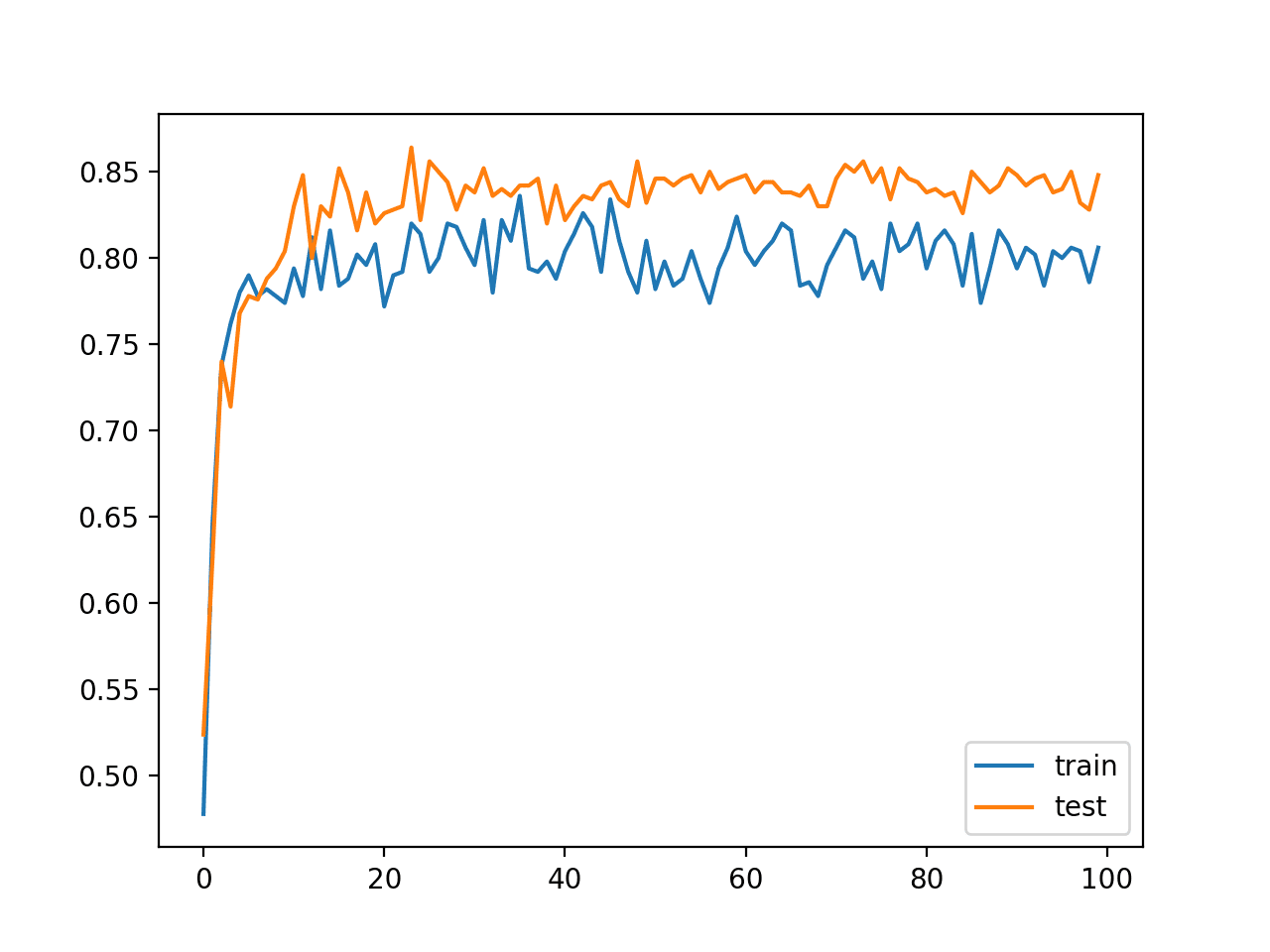
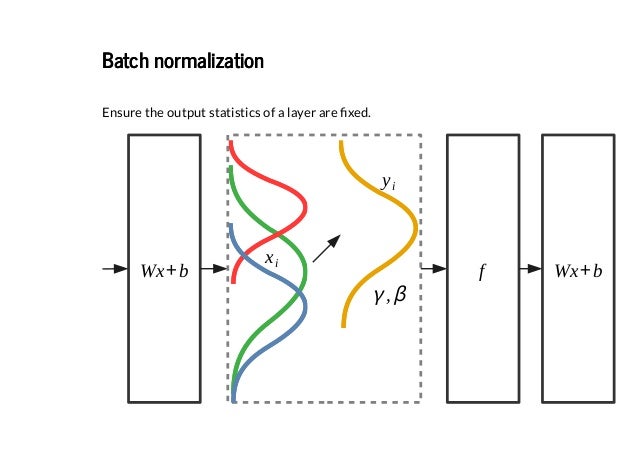
How to Accelerate Learning of Deep Neural Networks With Batch Normalization - MachineLearningMastery.com
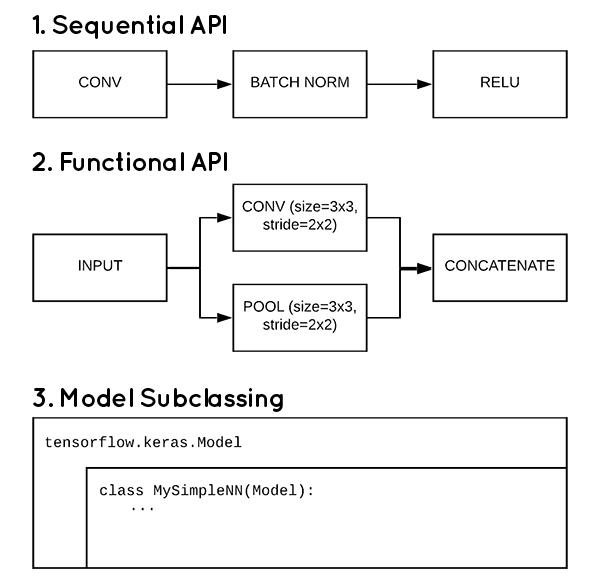
3 ways to create a Keras model with TensorFlow 2.0 (Sequential, Functional, and Model Subclassing) - PyImageSearch
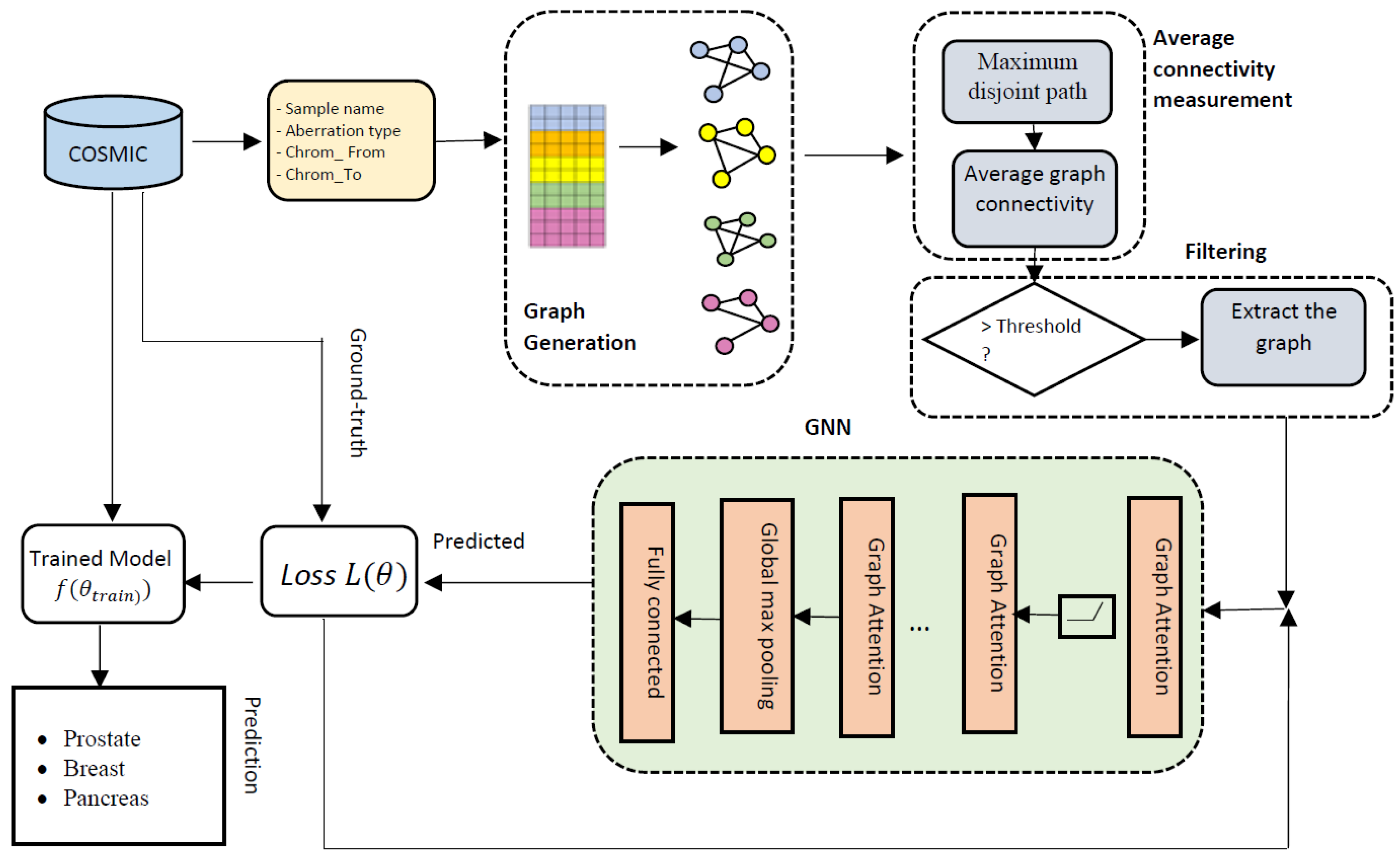
Cancers | Free Full-Text | GraphChrom: A Novel Graph-Based Framework for Cancer Classification Using Chromosomal Rearrangement Endpoints

python - Tensorflow tf.math.tanh properly scale network output without requiring large batches - Stack Overflow
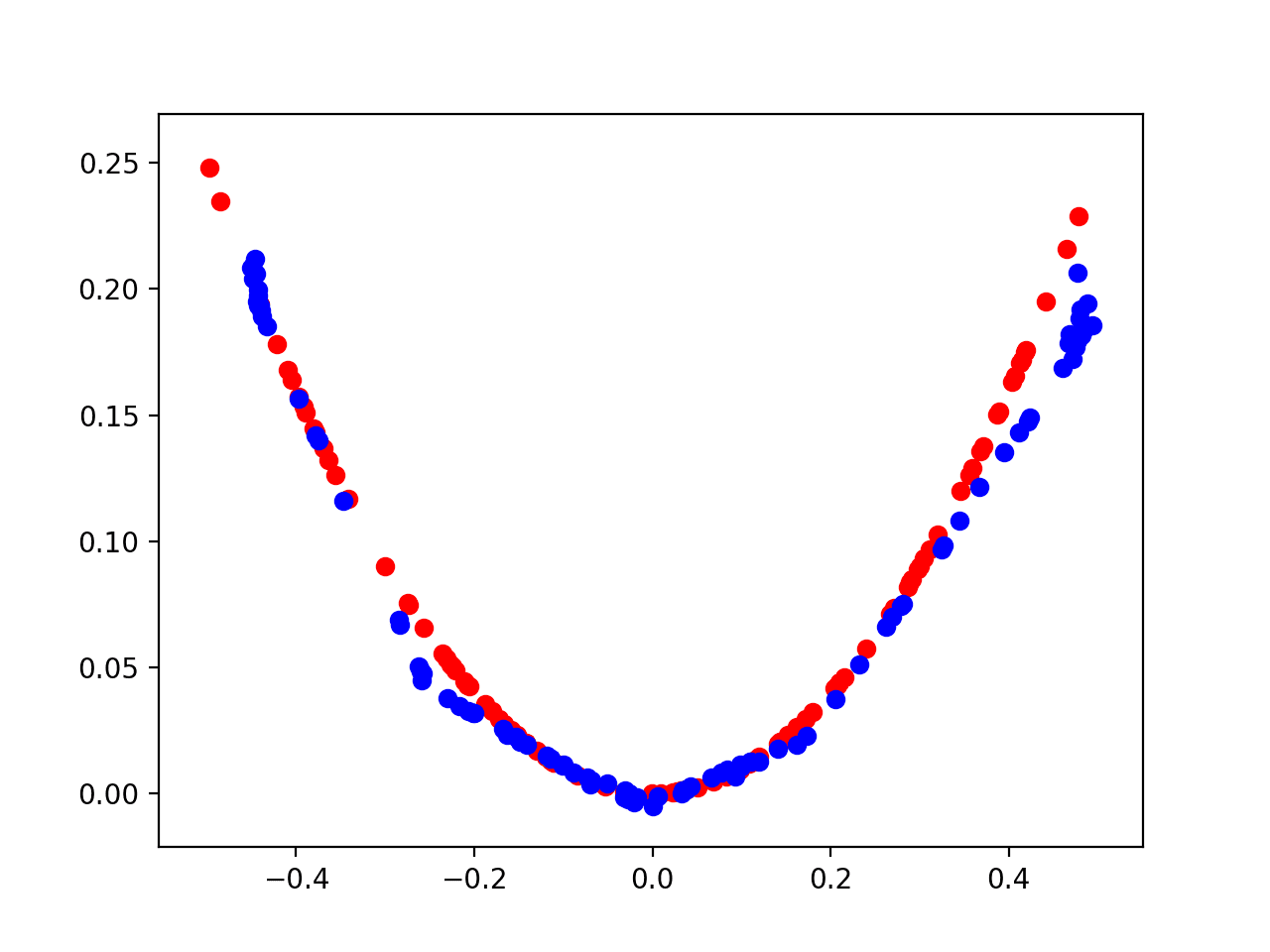
How to Develop a 1D Generative Adversarial Network From Scratch in Keras - MachineLearningMastery.com
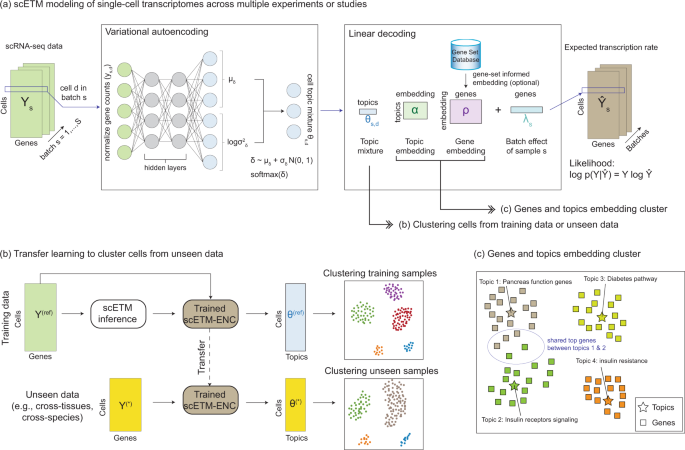
Learning interpretable cellular and gene signature embeddings from single-cell transcriptomic data | Nature Communications
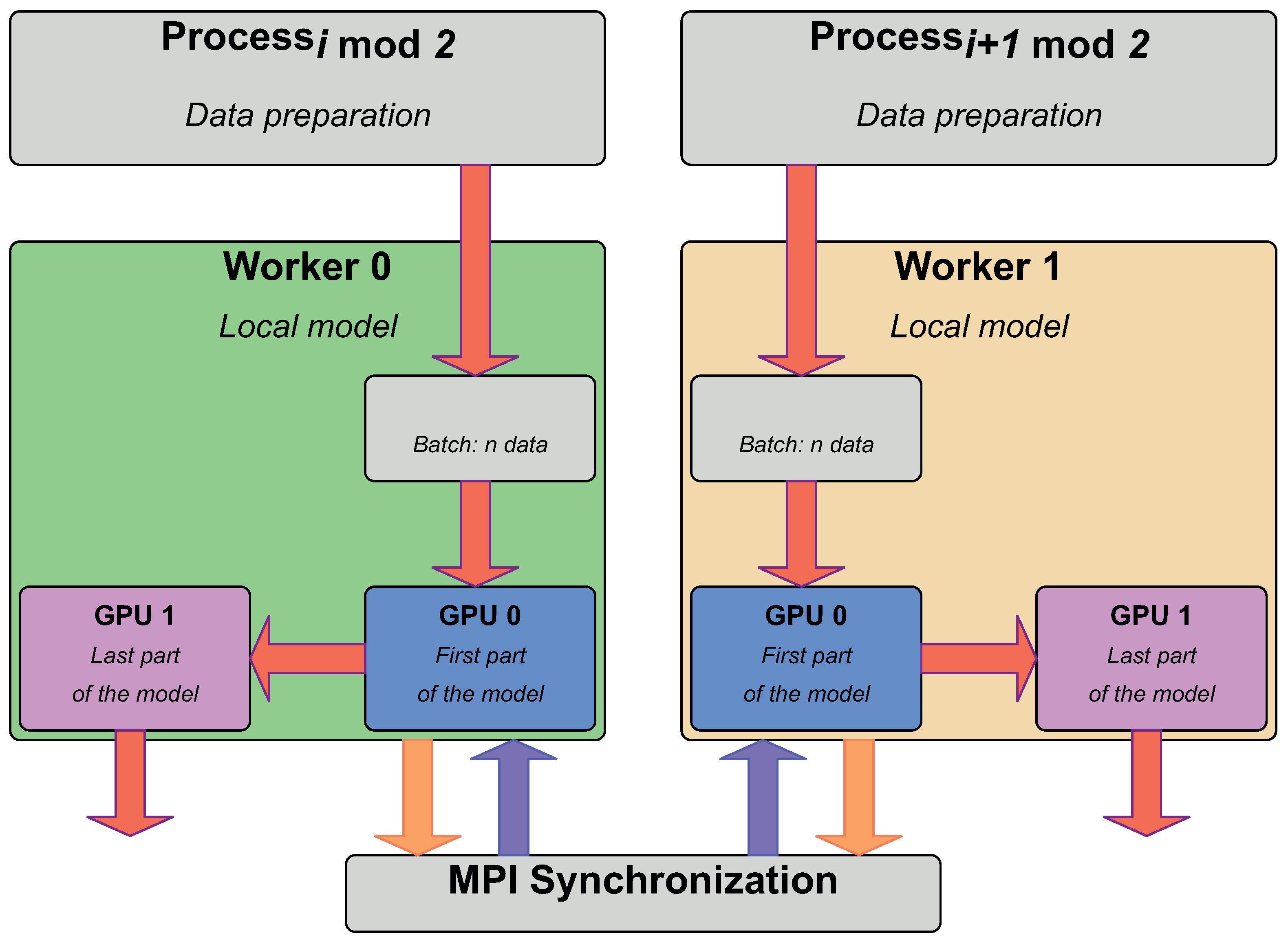
Electronics | Free Full-Text | Distributed Deep Learning: From Single-Node to Multi-Node Architecture
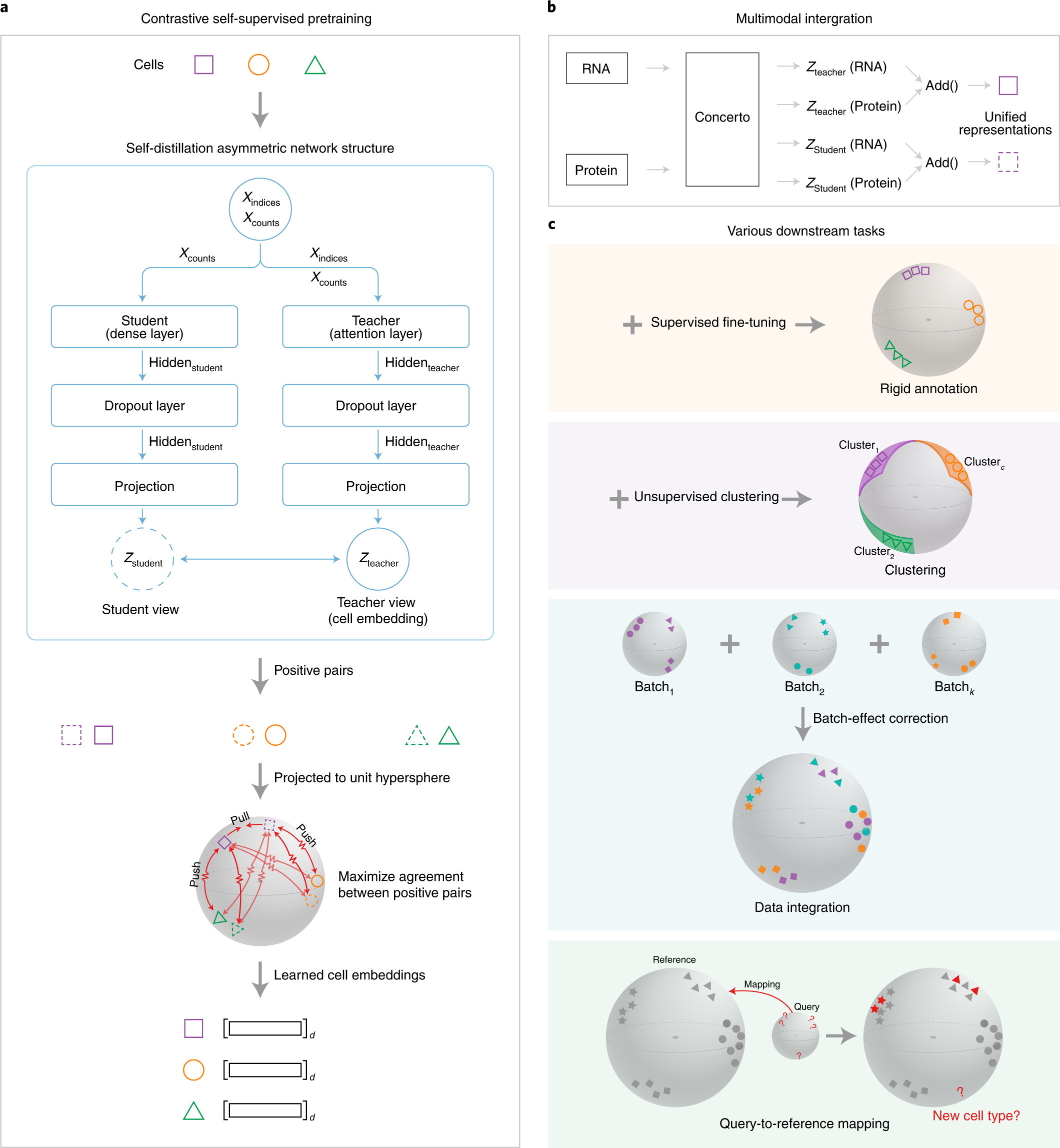
Contrastive learning enables rapid mapping to multimodal single-cell atlas of multimillion scale | Nature Machine Intelligence